APT1, but not APT2, codes for a functional adenine phosphoribosyltransferase in Saccharomyces cerevisiae
- PMID: 9864350
- PMCID: PMC103569
- DOI: 10.1128/JB.181.1.347-352.1999
APT1, but not APT2, codes for a functional adenine phosphoribosyltransferase in Saccharomyces cerevisiae
Abstract
The yeast Saccharomyces cerevisiae has two separate genes (APT1 and APT2) that encode two potentially different forms of adenine phosphoribosyltransferase (APRT). However, genetic analysis indicated that only APT1 could code for a complementing activity. Cloning and expression of both the APT1 and APT2 genes in Escherichia coli showed that although discrete proteins (APRT1 and APRT2) were made by these genes, only APRT1 had detectable APRT activity. Northern and Western blot analyses demonstrated that only APT1 was transcribed and translated under normal physiological conditions in yeast. Phylogenetic analysis revealed that APRT1 and APRT2 are evolutionary closely related and that they arise from a gene duplication event. We conclude that APT1 is the functional gene in S. cerevisiae and that APT2 is a pseudogene.
Figures

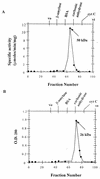



References
-
- Alfonzo J D. Ph.D. thesis. Bloomington, Ind: Indiana University; 1995.
-
- Alfonzo J D, Sahota A, Taylor M W. Purification and characterization of adenine phosphoribosyltransferase from Saccharomyces cerevisiae. Biochim Biophys Acta. 1997;1341:173–182. - PubMed
-
- Alfonzo J D, Sahota A, Deeley M C, Ranjekar P, Taylor M W. Cloning and characterization of the adenine phosphoribosyltransferase-encoding gene (APT1) from Saccharomyces cerevisiae. Gene. 1995;161:81–85. - PubMed
-
- Allen J B, Elledge S J. A family of vectors that facilitate transposon and insertional mutagenesis of cloned genes in yeast. Yeast. 1994;10:1267–1272. - PubMed
-
- Arnold W J. Adenine phosphoribosyltransferase. Methods Enzymol. 1978;51:568–574. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

