Visualization of protein-RNA interactions in cytoplasmic polyhedrosis virus
- PMID: 9882369
- PMCID: PMC103988
- DOI: 10.1128/JVI.73.2.1624-1629.1999
Visualization of protein-RNA interactions in cytoplasmic polyhedrosis virus
Abstract
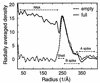
Unlike the multiple-shelled organization of other Reoviridae members, cytoplasmic polyhedrosis virus (CPV) has a single-shelled capsid. The three-dimensional structures of full and empty CPV by electron cryomicroscopy show identical outer shells but differ inside. The outer surface reveals a T=1 icosahedral shell decorated with spikes at its icosahedral vertices. The internal space of the empty CPV is unoccupied except for 12 mushroom-shaped densities attributed to the transcriptional enzyme complexes. The ordered double-stranded RNA inside the full capsid forms spherical shells spaced 25 A apart. The RNA-protein interactions suggest a mechanism for RNA transcription and release.
Figures



References
-
- Crowther R A. Procedures for three-dimensional reconstruction of spherical viruses by Fourier synthesis from electron micrographs. Phil Trans R Soc Lond B. 1971;261:221–230. - PubMed
-
- Dai R M, Wu A Z, Sun Y K. The protein subunits of the double-stranded RNA dependent RNA polymerase and methyltransferase of the cytoplasmic polyhedrosis virus of silkworm, Bombyx mori. Sci Sin Ser B (Chem Biol Agric Med Earth Sci) 1986;29:1267–1272. - PubMed
-
- Dryden K A, Farsetta D L, Wang G, Keegan J M, Fields B N, Baker T S, Nibert M L. Internal structures containing transcriptase-related proteins in top component particles of mammalian orthoreovirus. Virology. 1998;245:33–46. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

