Functional analyses of the promoters in the lantibiotic mutacin II biosynthetic locus in Streptococcus mutans
- PMID: 9925596
- PMCID: PMC91075
- DOI: 10.1128/AEM.65.2.652-658.1999
Functional analyses of the promoters in the lantibiotic mutacin II biosynthetic locus in Streptococcus mutans
Abstract
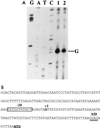
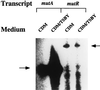
The lantibiotic bacteriocin mutacin II is produced by the group II Streptococcus mutans. The mutacin II biosynthetic locus consists of seven genes, mutR, -A, -M, -T, -F, -E, and -G, organized as two operons. The mutAMTFEG operon is transcribed from the mutA promoter 55 bp upstream of the translation start codon for MutA, while the mutR promoter is 76 bp upstream of the mutR structural gene. Expression of the mutA promoter is regulated by the components of the growth medium, while the mutR promoter activity does not seem to be affected by these conditions. Inactivation of mutR abolishes transcription of the mutA operon but does not affect its own promoter activity. The expressions of both mutA and mutR promoters are independent of the growth stage, while the production of mutacin II is only elevated at the early stationary phase. Taken together, these results suggest that expression of the mutacin operon is regulated by a complex system involving transcriptional and posttranscriptional or posttranslational controls.
Figures
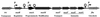
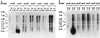



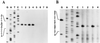
References
-
- Bibb M. The regulation of antibiotic production in Streptomyces coelicolor A3(2) Microbiology. 1996;142:1335–1344. - PubMed
-
- Caufield P W, Chen P, Qi F, Novak J. Presented at the 5th Conference on Streptococcal Genetics, Vichy, France. 1998.
-
- Chen P, Novak J, Qi F, Caufield P W. Presented at the Third International Workshop on Lantibiotics and Related Modified Antibiotic Peptides, Blaubeuren, Germany. 1998.
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

