Conversion of 3-chlorocatechol by various catechol 2,3-dioxygenases and sequence analysis of the chlorocatechol dioxygenase region of Pseudomonas putida GJ31
- PMID: 9973359
- PMCID: PMC93510
- DOI: 10.1128/JB.181.4.1309-1318.1999
Conversion of 3-chlorocatechol by various catechol 2,3-dioxygenases and sequence analysis of the chlorocatechol dioxygenase region of Pseudomonas putida GJ31
Abstract
Pseudomonas putida GJ31 contains an unusual catechol 2,3-dioxygenase that converts 3-chlorocatechol and 3-methylcatechol, which enables the organism to use both chloroaromatics and methylaromatics for growth. A 3.1-kb region of genomic DNA of strain GJ31 containing the gene for this chlorocatechol 2,3-dioxygenase (cbzE) was cloned and sequenced. The cbzE gene appeared to be plasmid localized and was found in a region that also harbors genes encoding a transposase, a ferredoxin that was homologous to XylT, an open reading frame with similarity to a protein of a meta-cleavage pathway with unknown function, and a 2-hydroxymuconic semialdehyde dehydrogenase. CbzE was most similar to catechol 2,3-dioxygenases of the 2.C subfamily of type 1 extradiol dioxygenases (L. D. Eltis and J. T. Bolin, J. Bacteriol. 178:5930-5937, 1996). The substrate range and turnover capacity with 3-chlorocatechol were determined for CbzE and four related catechol 2,3-dioxygenases. The results showed that CbzE was the only enzyme that could productively convert 3-chlorocatechol. Besides, CbzE was less susceptible to inactivation by methylated catechols. Hybrid enzymes that were made of CzbE and the catechol 2, 3-dioxygenase of P. putida UCC2 (TdnC) showed that the resistance of CbzE to suicide inactivation and its substrate specificity were mainly determined by the C-terminal region of the protein.
Figures
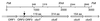



 , polypeptide fragments derived from TdnC. The amino acids of
CbzE and TdnC between which the fusions were made are indicated.
, polypeptide fragments derived from TdnC. The amino acids of
CbzE and TdnC between which the fusions were made are indicated.References
-
- Assinder S J, Williams P A. The TOL plasmids: determinants of the catabolism of toluene and the xylenes. Adv Microb Physiol. 1990;31:1–69. - PubMed
-
- Ausubel F M, Brent R, Kingston R E, Moore D D, Seidman J G, Smith J A, Struhl K. Current protocols in molecular biology. New York, N.Y: Green Publishing Associates, Inc.; 1994.
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

