Large-scale engineering of the Corynebacterium glutamicum genome
- PMID: 15933044
- PMCID: PMC1151864
- DOI: 10.1128/AEM.71.6.3369-3372.2005
Large-scale engineering of the Corynebacterium glutamicum genome
Abstract
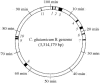
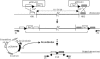
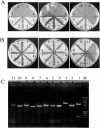
The engineering of Corynebacterium glutamicum is important for enhanced production of biochemicals. To construct an improved C. glutamicum genome, we developed a precise genome excision method based on the Cre/loxP recombination system and successfully deleted 11 distinct genomic regions identified by comparative analysis of C. glutamicum genomes. Despite the loss of several predicted open reading frames, the mutant cells exhibited normal growth under standard laboratory conditions. With a total of 250 kb (7.5% of the genome), the 11 genomic regions were loaded with cryptic prophages, transposons, and genes of unknown function which were dispensable for cell growth, indicating recent horizontal acquisitions to the genome. This provides an interesting background for functional genomic studies and can be used in the improvement of cell traits.
Figures
References
-
- Campo, N., M. J. Dias, M. L. Daveran-Mingot, P. Ritzenthaler, and P. Le Bourgeois. 2004. Chromosomal constraints in Gram-positive bacteria revealed by artificial inversions. Mol. Microbiol. 51:511-522. - PubMed
-
- Casjens, S. 2003. Prophages and bacterial genomics: what have we learned so far? Mol. Microbiol. 49:277-300. - PubMed
-
- Eikmanns, B. J., E. Kleinertz, W. Liebl, and H. Sahm. 1991. A family of Corynebacterium glutamicum/Escherichia coli shuttle vectors for cloning, controlled gene expression, and promoter probing. Gene 102:93-98. - PubMed
-
- Hacker, J., G. Blum-Oehler, I. Muhldorfer, and H. Tschape. 1997. Pathogenicity islands of virulent bacteria: structure, function and impact on microbial evolution. Mol. Microbiol. 23:1089-1097. - PubMed
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

