A Sabin 3-derived poliovirus recombinant contained a sequence homologous with indigenous human enterovirus species C in the viral polymerase coding region
- PMID: 16188967
- PMCID: PMC1235834
- DOI: 10.1128/JVI.79.20.12650-12657.2005
A Sabin 3-derived poliovirus recombinant contained a sequence homologous with indigenous human enterovirus species C in the viral polymerase coding region
Abstract
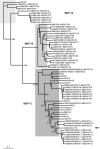
Outbreaks of poliomyelitis caused by circulating vaccine-derived polioviruses (cVDPVs) have been reported in areas where indigenous wild polioviruses (PVs) were eliminated by vaccination. Most of these cVDPVs contained unidentified sequences in the nonstructural protein coding region which were considered to be derived from human enterovirus species C (HEV-C) by recombination. In this study, we report isolation of a Sabin 3-derived PV recombinant (Cambodia-02) from an acute flaccid paralysis (AFP) case in Cambodia in 2002. We attempted to identify the putative recombination counterpart of Cambodia-02 by sequence analysis of nonpolio enterovirus isolates from AFP cases in Cambodia from 1999 to 2003. Based on the previously estimated evolution rates of PVs, the recombination event resulting in Cambodia-02 was estimated to have occurred within 6 months after the administration of oral PV vaccine (99.3% nucleotide identity in VP1 region). The 2BC and the 3D(pol) coding regions of Cambodia-02 were grouped into the genetic cluster of indigenous coxsackie A virus type 17 (CAV17) (the highest [87.1%] nucleotide identity) and the cluster of indigenous CAV13-CAV18 (the highest [94.9%] nucleotide identity) by the phylogenic analysis of the HEV-C isolates in 2002, respectively. CAV13-CAV18 and CAV17 were the dominant HEV-C serotypes in 2002 but not in 2001 and in 2003. We found a putative recombination between CAV13-CAV18 and CAV17 in the 3CD(pro) coding region of a CAV17 isolate. These results suggested that a part of the 3D(pol) coding region of PV3(Cambodia-02) was derived from a HEV-C strain genetically related to indigenous CAV13-CAV18 strains in 2002 in Cambodia.
Figures



References
-
- Alexander, J. P., Jr., H. E. Gary, Jr., and M. A. Pallansch. 1997. Duration of poliovirus excretion and its implications for acute flaccid paralysis surveillance: a review of the literature. J. Infect. Dis. 175:S176-S182. - PubMed
-
- Bellmunt, A., G. May, R. Zell, P. Pring-Akerblom, W. Verhagen, and A. Heim. 1999. Evolution of poliovirus type I during 5.5 years of prolonged enteral replication in an immunodeficient patient. Virology 265:178-184. - PubMed
-
- Caro, V., S. Guillot, F. Delpeyroux, and R. Crainic. 2001. Molecular strategy for ‘serotyping’ of human enteroviruses. J. Gen. Virol. 82:79-91. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Medical
Research Materials

