Hypoxia-induced gene expression profiling in the euryoxic fish Gillichthys mirabilis
- PMID: 11172064
- PMCID: PMC29370
- DOI: 10.1073/pnas.98.4.1993
Hypoxia-induced gene expression profiling in the euryoxic fish Gillichthys mirabilis
Abstract
Hypoxia is important in both biomedical and environmental contexts and necessitates rapid adaptive changes in metabolic organization. Mammals, as air breathers, have a limited capacity to withstand sustained exposure to hypoxia. By contrast, some aquatic animals, such as certain fishes, are routinely exposed and resistant to severe environmental hypoxia. Understanding the changes in gene expression in fishes exposed to hypoxic stress could reveal novel mechanisms of tolerance that may shed new light on hypoxia and ischemia in higher vertebrates. Using cDNA microarrays, we have studied gene expression in a hypoxia-tolerant burrow-dwelling goby fish, Gillichthys mirabilis. We show that a coherent picture of a complex transcriptional response can be generated for a nonmodel organism for which sequence data were unavailable. We demonstrate that: (i) although certain shifts in gene expression mirror changes in mammals, novel genes are differentially expressed in fish; and (ii) tissue-specific patterns of expression reflect the different metabolic roles of tissues during hypoxia.
Figures
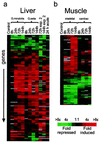

References
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources

