Comparison of diversities and compositions of bacterial populations inhabiting natural forest soils
- PMID: 15345382
- PMCID: PMC520910
- DOI: 10.1128/AEM.70.9.5057-5065.2004
Comparison of diversities and compositions of bacterial populations inhabiting natural forest soils
Abstract
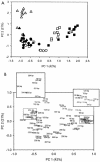
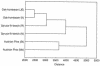
The diversity and composition of soil bacterial communities were compared among six Austrian natural forests, including oak-hornbeam, spruce-fir-beech, and Austrian pine forests, using terminal restriction fragment length polymorphism (T-RFLP, or TRF) analysis and sequence analysis of 16S rRNA genes. The forests studied differ greatly in soil chemical characteristics, microbial biomass, and nutrient turnover rates. The aim of this study was to relate these differences to the composition of the bacterial communities inhabiting the individual forest soils. Both TRF profiling and clone sequence analysis revealed that the bacterial communities in soils under Austrian pine forests, representing azonal forest types, were distinct from those in soils under zonal oak-hornbeam and spruce-fir-beech forests, which were more similar in community composition. Clones derived from an Austrian pine forest soil were mostly affiliated with high-G+C gram-positive bacteria (49%), followed by members of the alpha-Proteobacteria (20%) and the Holophaga/Acidobacterium group (12%). Clones in libraries from oak-hornbeam and spruce-fir-beech forest soils were mainly related to the Holophaga/Acidobacterium group (28 and 35%), followed by members of the Verrucomicrobia (24%) and the alpha-Proteobacteria (27%), respectively. The soil bacterial communities in forests with distinct vegetational and soil chemical properties appeared to be well differentiated based on 16S rRNA gene phylogeny. In particular, the outstanding position of the Austrian pine forests, which are determined by specific soil conditions, was reflected in the bacterial community composition.
Figures

References
-
- Axelrood, P. E., M. L. Chow, C. C. Radomski, J. M. McDermott, and J. Davies. 2002. Molecular characterization of bacterial diversity from British Columbia forest soils subjected to disturbance. Can. J. Microbiol. 48:655-674. - PubMed
-
- Chow, M. L., C. C. Radomski, J. M. McDermott, J. Davies, and P. E. Axelrood. 2002. Molecular characterisation of bacterial diversity in lodgepole pine (Pinus contorta) rhizosphere soils from British Columbia forest soils differing in disturbance and geographic source. FEMS Microbiol. Ecol. 42:347-357. - PubMed
-
- Cole, J. R., B. Chai, T. L. Marsh, R. J. Farris, Q. Wang, S. A. Kulam, S. Chandra, D. M. McGarrell, T. M. Schmidt, G. M. Garrity, and J. M. Tiedje. 2003. The Ribosomal Database Project (RDP-II): previewing a new autoaligner that allows regular updates and the new prokaryotic taxonomy. Nucleic Acids Res. 31:442-443. - PMC - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

