Simian T-cell leukemia virus (STLV) infection in wild primate populations in Cameroon: evidence for dual STLV type 1 and type 3 infection in agile mangabeys (Cercocebus agilis)
- PMID: 15078952
- PMCID: PMC387697
- DOI: 10.1128/jvi.78.9.4700-4709.2004
Simian T-cell leukemia virus (STLV) infection in wild primate populations in Cameroon: evidence for dual STLV type 1 and type 3 infection in agile mangabeys (Cercocebus agilis)
Abstract
Three types of human T-cell leukemia virus (HTLV)-simian T-cell leukemia virus (STLV) (collectively called primate T-cell leukemia viruses [PTLVs]) have been characterized, with evidence for zoonotic origin from primates for HTLV type 1 (HTLV-1) and HTLV-2 in Africa. To assess human exposure to STLVs in western Central Africa, we screened for STLV infection in primates hunted in the rain forests of Cameroon. Blood was obtained from 524 animals representing 18 different species. All the animals were wild caught between 1999 and 2002; 328 animals were sampled as bush meat and 196 were pets. Overall, 59 (11.2%) of the primates had antibodies cross-reacting with HTLV-1 and/or HTLV-2 antigens; HTLV-1 infection was confirmed in 37 animals, HTLV-2 infection was confirmed in 9, dual HTLV-1 and HTLV-2 infection was confirmed in 10, and results for 3 animals were indeterminate. Prevalences of infection were significantly lower in pets than in bush meat, 1.5 versus 17.0%, respectively. Discriminatory PCRs identified STLV-1, STLV-3, and STLV-1 and STLV-3 in HTLV-1-, HTLV-2-, and HTLV-1- and HTLV-2-cross-reactive samples, respectively. We identified for the first time STLV-1 sequences in mustached monkeys (Cercopithecus cephus), talapoins (Miopithecus ogouensis), and gorillas (Gorilla gorilla) and confirmed STLV-1 infection in mandrills, African green monkeys, agile mangabeys, and crested mona and greater spot-nosed monkeys. STLV-1 long terminal repeat (LTR) and env sequences revealed that the strains belonged to different PTLV-1 subtypes. A high prevalence of PTLV infection was observed among agile mangabeys (Cercocebus agilis); 89% of bush meat was infected with STLV. Cocirculation of STLV-1 and STLV-3 and STLV-1-STLV-3 coinfections were identified among the agile mangabeys. Phylogenetic analyses of partial LTR sequences indicated that the agile mangabey STLV-3 strains were more related to the STLV-3 CTO604 strain isolated from a red-capped mangabey (Cercocebus torquatus) from Cameroon than to the STLV-3 PH969 strain from an Eritrean baboon or the PPA-F3 strain from a baboon in Senegal. Our study documents for the first time that (i) a substantial proportion of wild-living monkeys in Cameroon is STLV infected, (ii) STLV-1 and STLV-3 cocirculate in the same primate species, (iii) coinfection with STLV-1 and STLV-3 occurs in agile mangabeys, and (iv) humans are exposed to different STLV-1 and STLV-3 subtypes through handling primates as bush meat.
Figures




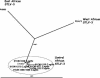
References
-
- Asibey, E. O. 1974. Wildlife as a source of protein in Africa south of the Sahara. Biol. Conserv. 6:32-39.
-
- Auzel, P., and R. Hardin. 2000. Colonial history, concessionary politics, and collaborative management of Equatorial African rain forests, p. 21-38. In M. Bakarr, G. Da Fonseca, W. Konstant, R. Mittermeier, and K. Painemilla (ed.), Hunting and bushmeat utilization in the African rain forest. Conservation International, Washington, D.C.
-
- Bennett, E. L., and J. G. Robinson. 2000. Hunting for the snark, p. 1-9. In J. G. Robinson, and E. L. Bennett (ed.), Hunting for sustainability in tropical forests. Columbia University Press, New York, N.Y.
-
- Delaporte, E., A. Dupont, M. Peeters, R. Josse, M. Merlin, D. Schrijvers, B. Hamono, L. Bedjabaga, H. Cheringou, F. Boyer, et al. 1988. Epidemiology of HTLV-I in Gabon (Western Equatorial Africa). Int. J. Cancer 42:687-689. - PubMed
-
- Delaporte, E., M. Peeters, J. P. Durand, A. Dupont, D. Schrijvers, L. Bedjabaga, C. Honore, S. Ossari, A. Trebucq, R. Josse, et al. 1989. Seroepidemiological survey of HTLV-I infection among randomized populations of western central African countries. J. Acquir. Immune Defic. Syndr. 2:410-413. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

