Population-based genetic and evolutionary analysis of Chlamydia trachomatis urogenital strain variation in the United States
- PMID: 15060049
- PMCID: PMC412158
- DOI: 10.1128/JB.186.8.2457-2465.2004
Population-based genetic and evolutionary analysis of Chlamydia trachomatis urogenital strain variation in the United States
Abstract
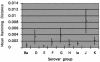
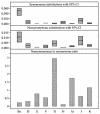
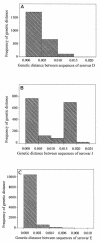
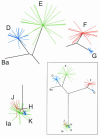
Chlamydia trachomatis is a major cause of ocular and sexually transmitted diseases worldwide. While much of our knowledge about its genetic diversity comes from serotyping or ompA genotyping, no quantitative assessment of genetic diversity within serotypes has been performed. To accomplish this, 507 urogenital samples from a multicenter U.S. study were analyzed by phylogenetic and statistical modeling. No B, Da, or I serotypes were represented. Based on our analyses, all but one previous urogenital B serotype was identified as Ba. This, coupled with the lack of B serotypes in our population, suggests that B has specific tropism for ocular mucosa. We identified a Ba/D recombinant (putative crossover nucleotide 477; P < 0.0001) similar to a B/D mosaic we described previously from an African trachoma patient. Computational analyses of the Ba/D recombinant indicated that upstream changes were less important for tissue tropism than downstream incorporation of the D sequence. Since most serotypes had nonsynonymous/synonymous ratios of <1.0, the major outer membrane protein, encoded by ompA, has many functional constraints and is under purifying selection. Surprisingly, all serotype groups except for J had a unimodal population structure indicating rapid clonal expansion. Of the groups with a unimodal structure, E and Ia and, to a lesser extent, G and K were prevalent, had infrequent incorporation of mutations, and, compared to other groups, had a relatively greater degree of diversifying selection, consistent with a selective sweep of mutations within these groups. Collectively, these data suggest a diverse evolutionary strategy for different serogroups of the organism.
Figures

References
-
- Allen, J. E., R. M. Locksley, and R. S. Stephens. 1991. A single peptide from the major outer membrane protein of Chlamydia trachomatis elicits T cell help for the production of antibodies to protective determinants. J. Immunol. 147:674-679. - PubMed
-
- Bandea, C. I., K. Kubota, T. M. Brown, P. H. Kilmarx, V. Bhullar, S. Yanpaisarn, P. Chaisilwattana, W. Siriwasin, and C. M. Black. 2001. Typing of Chlamydia trachomatis strains from urine samples by amplification and sequencing the major outer membrane protein gene (omp1). Sex. Transm. Infect. 77:419-422. - PMC - PubMed
-
- Barnes, R. C., A. M. Rompalo, and W. E. Stamm. 1987. Comparison of Chlamydia trachomatis serovars causing rectal and cervical infections. J. Infect. Dis. 156:953-958. - PubMed
-
- Batteiger, B. E., J. Fraiz, W. J. Newhall, B. P. Katz, and R. B. Jones. 1989. Association of recurrent chlamydial infection with gonorrhea. J. Infect. Dis. 159:661-669. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials

