Analysis of the Basidiomycete Coprinopsis cinerea reveals conservation of the core meiotic expression program over half a billion years of evolution
- PMID: 20885784
- PMCID: PMC2944786
- DOI: 10.1371/journal.pgen.1001135
Analysis of the Basidiomycete Coprinopsis cinerea reveals conservation of the core meiotic expression program over half a billion years of evolution
Abstract
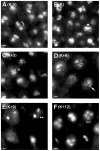
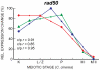
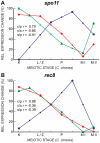
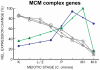
Coprinopsis cinerea (also known as Coprinus cinereus) is a multicellular basidiomycete mushroom particularly suited to the study of meiosis due to its synchronous meiotic development and prolonged prophase. We examined the 15-hour meiotic transcriptional program of C. cinerea, encompassing time points prior to haploid nuclear fusion though tetrad formation, using a 70-mer oligonucleotide microarray. As with other organisms, a large proportion (∼20%) of genes are differentially regulated during this developmental process, with successive waves of transcription apparent in nine transcriptional clusters, including one enriched for meiotic functions. C. cinerea and the fungi Saccharomyces cerevisiae and Schizosaccharomyces pombe diverged ∼500-900 million years ago, permitting a comparison of transcriptional programs across a broad evolutionary time scale. Previous studies of S. cerevisiae and S. pombe compared genes that were induced upon entry into meiosis; inclusion of C. cinerea data indicates that meiotic genes are more conserved in their patterns of induction across species than genes not known to be meiotic. In addition, we found that meiotic genes are significantly more conserved in their transcript profiles than genes not known to be meiotic, which indicates a remarkable conservation of the meiotic process across evolutionarily distant organisms. Overall, meiotic function genes are more conserved in both induction and transcript profile than genes not known to be meiotic. However, of 50 meiotic function genes that were co-induced in all three species, 41 transcript profiles were well-correlated in at least two of the three species, but only a single gene (rad50) exhibited coordinated induction and well-correlated transcript profiles in all three species, indicating that co-induction does not necessarily predict correlated expression or vice versa. Differences may reflect differences in meiotic mechanisms or new roles for paralogs. Similarities in induction, transcript profiles, or both, should contribute to gene discovery for orthologs without currently characterized meiotic roles.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures




References
-
- Redhead SA, Vilgalys R, Moncalvo JM, Johnson J, Hopple JS. Coprinus Pers. and the disposition of Coprinus species sensu lato. Taxon. 2001;50:203–241.
-
- Raju NB, Lu BC. Meiosis in Coprinus. 3: Timing of meiotic events in C. lagopus (sensu Buller). Canadian Journal of Botany. 1970;48:2183–2186.
-
- Page SL, Hawley RS. The genetics and molecular biology of the synaptonemal complex. Annual Review of Cell and Developmental Biology. 2004;20:525–558. - PubMed
-
- Henderson KA, Keeney S. Synaptonemal complex formation: where does it start? Bioessays. 2005;27:995–998. - PubMed
-
- Li WX, Ma H. Double-stranded DNA breaks and gene functions in recombination and meiosis. Cell Research. 2006;16:402–412. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

