Circadian transcriptional regulation by the posttranslational oscillator without de novo clock gene expression in Synechococcus
- PMID: 21896749
- PMCID: PMC3174641
- DOI: 10.1073/pnas.1019612108
Circadian transcriptional regulation by the posttranslational oscillator without de novo clock gene expression in Synechococcus
Abstract
Circadian rhythms are a fundamental property of most organisms, from cyanobacteria to humans. In the unicellular obligately photoautotrophic cyanobacterium Synechococcus elongatus PCC 7942, essentially all promoter activities are controlled by the KaiABC-based clock under continuous light conditions. When Synechococcus cells are transferred from the light to continuous dark (DD) conditions, the expression of most genes, including the clock genes kaiA and kaiBC, is rapidly down-regulated, whereas the KaiC phosphorylation cycle persists. Therefore, we speculated that the posttranslational oscillator might not drive the transcriptional circadian output without de novo expression of the kai genes. Here we show that the cyanobacterial clock regulates the transcriptional output even in the dark. The expression of a subset of genes in the genomes of cells grown in the dark was dramatically affected by kaiABC nullification, and the magnitude of dark induction was dependent on the time at which the cells were transferred from the light to the dark. Moreover, under DD conditions, the expression of some dark-induced gene transcripts exhibited temperature-compensated damped oscillations, which were nullified in kaiABC-null strains and were affected by a kaiC period mutation. These results indicate that the Kai protein-based posttranslational oscillator can drive the circadian transcriptional output even without the de novo expression of the clock genes.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

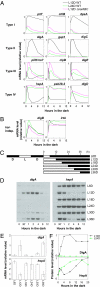


References
-
- Ishiura M, et al. Expression of a gene cluster kaiABC as a circadian feedback process in cyanobacteria. Science. 1998;281:1519–1523. - PubMed
-
- Tomita J, Nakajima M, Kondo T, Iwasaki H. No transcription-translation feedback in circadian rhythm of KaiC phosphorylation. Science. 2005;307:251–254. - PubMed
-
- Nakajima M, et al. Reconstitution of circadian oscillation of cyanobacterial KaiC phosphorylation in vitro. Science. 2005;308:414–415. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

