Human respiratory organoids sustained reproducible propagation of human rhinovirus C and elucidation of virus-host interaction
- PMID: 39738014
- PMCID: PMC11686133
- DOI: 10.1038/s41467-024-55076-2
Human respiratory organoids sustained reproducible propagation of human rhinovirus C and elucidation of virus-host interaction
Abstract
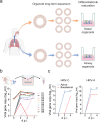
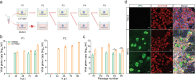
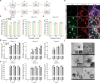
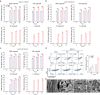
The lack of a robust system to reproducibly propagate HRV-C, a family of viruses refractory to cultivation in standard cell lines, has substantially hindered our understanding of this common respiratory pathogen. We sought to develop an organoid-based system to reproducibly propagate HRV-C, and characterize virus-host interaction using respiratory organoids. We demonstrate that airway organoids sustain serial virus passage with the aid of CYT387-mediated immunosuppression, whereas nasal organoids that more closely simulate the upper airway achieve this without any intervention. Nasal organoids are more susceptible to HRV-C than airway organoids. Intriguingly, upon HRV-C infection, we observe an innate immune response that is stronger in airway organoids than in nasal organoids, which is reproduced in a Poly(I:C) stimulation assay. Treatment with α-CDHR3 and antivirals significantly reduces HRV-C viral growth in airway and nasal organoids. Additionally, an organoid-based immunofluorescence assay is established to titrate HRV-C infectious particles. Collectively, we develop an organoid-based system to reproducibly propagate the poorly cultivable HRV-C, followed by a comprehensive characterization of HRV-C infection and innate immunity in physiologically active respiratory organoids. The organoid-based HRV-C infection model can be extended for developing antiviral strategies. More importantly, our study has opened an avenue for propagating and studying other uncultivable human and animal viruses.
© 2024. The Author(s).
Conflict of interest statement
Competing interests: J.Z., K.Y.Y., H.Clevers, C.L., and M.C.C. are listed as inventors on the patent of airway organoids (Patent No: ZL 2019 8 0037552.0), and nasal organoids (US 63/358,795). J.Z. is the founder of BiomOrgan Ltd. All other authors declare no competing interests.
Figures


References
-
- To, K. K. W., Yip, C. C. Y. & Yuen, K.-Y. Rhinovirus – from bench to bedside. J. Formos. Med. Assoc.116, 496–504 (2017). - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
Associated data
- Actions
Grants and funding
- CID-HKU1-3/Food and Health Bureau of the Government of the Hong Kong Special Administrative Region | Health and Medical Research Fund (HMRF)
- COVID1903010-project 11/Food and Health Bureau of the Government of the Hong Kong Special Administrative Region | Health and Medical Research Fund (HMRF)
- 17105420/Research Grants Council, University Grants Committee (RGC, UGC)
- 17113724/Research Grants Council, University Grants Committee (RGC, UGC)
- C7042-21G/Research Grants Council, University Grants Committee (RGC, UGC)
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

