Diverse transcription factor binding features revealed by genome-wide ChIP-seq in C. elegans
- PMID: 21177963
- PMCID: PMC3032928
- DOI: 10.1101/gr.114587.110
Diverse transcription factor binding features revealed by genome-wide ChIP-seq in C. elegans
Abstract
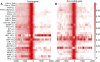
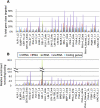
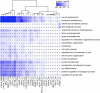
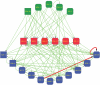
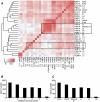
Regulation of gene expression by sequence-specific transcription factors is central to developmental programs and depends on the binding of transcription factors with target sites in the genome. To date, most such analyses in Caenorhabditis elegans have focused on the interactions between a single transcription factor with one or a few select target genes. As part of the modENCODE Consortium, we have used chromatin immunoprecipitation coupled with high-throughput DNA sequencing (ChIP-seq) to determine the genome-wide binding sites of 22 transcription factors (ALR-1, BLMP-1, CEH-14, CEH-30, EGL-27, EGL-5, ELT-3, EOR-1, GEI-11, HLH-1, LIN-11, LIN-13, LIN-15B, LIN-39, MAB-5, MDL-1, MEP-1, PES-1, PHA-4, PQM-1, SKN-1, and UNC-130) at diverse developmental stages. For each factor we determined candidate gene targets, both coding and non-coding. The typical binding sites of almost all factors are within a few hundred nucleotides of the transcript start site. Most factors target a mixture of coding and non-coding target genes, although one factor preferentially binds to non-coding RNA genes. We built a regulatory network among the 22 factors to determine their functional relationships to each other and found that some factors appear to act preferentially as regulators and others as target genes. Examination of the binding targets of three related HOX factors--LIN-39, MAB-5, and EGL-5--indicates that these factors regulate genes involved in cellular migration, neuronal function, and vulval differentiation, consistent with their known roles in these developmental processes. Ultimately, the comprehensive mapping of transcription factor binding sites will identify features of transcriptional networks that regulate C. elegans developmental processes.
Figures
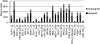

References
-
- Ao W, Gaudet J, Kent WJ, Muttumu S, Mango SE 2004. Environmentally induced foregut remodeling by PHA-4/FoxA and DAF-12/NHR. Science 305: 1743–1746 - PubMed
-
- Azzaria M, Goszczynski B, Chung MA, Kalb JM, McGhee JD 1996. A fork head/HNF-3 homolog expressed in the pharynx and intestine of the Caenorhabditis elegans embryo. Dev Biol 178: 289–303 - PubMed
-
- Bailey TL, Elkan C 1994. Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc Int Conf Intell Syst Mol Biol 2: 28–36 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous
