HIF1A employs CDK8-mediator to stimulate RNAPII elongation in response to hypoxia
- PMID: 23746844
- PMCID: PMC3681429
- DOI: 10.1016/j.cell.2013.04.048
HIF1A employs CDK8-mediator to stimulate RNAPII elongation in response to hypoxia
Abstract
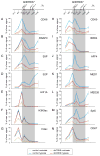
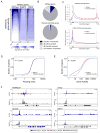
The transcription factor HIF1A is a key mediator of the cellular response to hypoxia. Despite the importance of HIF1A in homeostasis and various pathologies, little is known about how it regulates RNA polymerase II (RNAPII). We report here that HIF1A employs a specific variant of the Mediator complex to stimulate RNAPII elongation. The Mediator-associated kinase CDK8, but not the paralog CDK19, is required for induction of many HIF1A target genes. HIF1A induces binding of CDK8-Mediator and the super elongation complex (SEC), containing AFF4 and CDK9, to alleviate RNAPII pausing. CDK8 is dispensable for HIF1A chromatin binding and histone acetylation, but it is essential for binding of SEC and RNAPII elongation. Global analysis of active RNAPII reveals that hypoxia-inducible genes are paused and active prior to their induction. Our results provide a mechanistic link between HIF1A and CDK8, two potent oncogenes, in the cellular response to hypoxia.
Copyright © 2013 Elsevier Inc. All rights reserved.
Figures




References
-
- Adler AS, McCleland ML, Truong T, Lau S, Modrusan Z, Soukup TM, Roose-Girma M, Blackwood EM, Firestein R. CDK8 maintains tumor de-differentiation and embryonic stem cell pluripotency. Cancer Res. 2012;72:2129–2139. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

