Circadian patterns of gene expression in the human brain and disruption in major depressive disorder
- PMID: 23671070
- PMCID: PMC3683716
- DOI: 10.1073/pnas.1305814110
Circadian patterns of gene expression in the human brain and disruption in major depressive disorder
Abstract
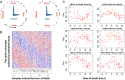
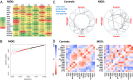
A cardinal symptom of major depressive disorder (MDD) is the disruption of circadian patterns. However, to date, there is no direct evidence of circadian clock dysregulation in the brains of patients who have MDD. Circadian rhythmicity of gene expression has been observed in animals and peripheral human tissues, but its presence and variability in the human brain were difficult to characterize. Here, we applied time-of-death analysis to gene expression data from high-quality postmortem brains, examining 24-h cyclic patterns in six cortical and limbic regions of 55 subjects with no history of psychiatric or neurological illnesses ("controls") and 34 patients with MDD. Our dataset covered ~12,000 transcripts in the dorsolateral prefrontal cortex, anterior cingulate cortex, hippocampus, amygdala, nucleus accumbens, and cerebellum. Several hundred transcripts in each region showed 24-h cyclic patterns in controls, and >100 transcripts exhibited consistent rhythmicity and phase synchrony across regions. Among the top-ranked rhythmic genes were the canonical clock genes BMAL1(ARNTL), PER1-2-3, NR1D1(REV-ERBa), DBP, BHLHE40 (DEC1), and BHLHE41(DEC2). The phasing of known circadian genes was consistent with data derived from other diurnal mammals. Cyclic patterns were much weaker in the brains of patients with MDD due to shifted peak timing and potentially disrupted phase relationships between individual circadian genes. This transcriptome-wide analysis of the human brain demonstrates a rhythmic rise and fall of gene expression in regions outside of the suprachiasmatic nucleus in control subjects. The description of its breakdown in MDD suggests potentially important molecular targets for treatment of mood disorders.
Keywords: circadian rhythms; depression; microarray.
Conflict of interest statement
Conflict of interest statement: The authors are members of the Pritzker Neuropsychiatric Disorders Research Consortium, which is supported by Pritzker Neuropsychiatric Disorders Research Fund, LLC. A shared intellectual property agreement exists between the academic and philanthropic entities of the consortium. The Pritzker Neuropsychiatric Disorders Research Fund had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Figures


References
-
- DeCoursey PJ. The behavioral ecology and evolution of biological timing systems. In: Dunlap JC, Loros JJ, Decoursey PJ, editors. Chronobiology: Biological Timekeeping. Sunderland, MA: Sinauer; 2004. pp. 26–65.
-
- Yamazaki S, et al. Resetting central and peripheral circadian oscillators in transgenic rats. Science. 2000;288(5466):682–685. - PubMed
-
- Akhtar RA, et al. Circadian cycling of the mouse liver transcriptome, as revealed by cDNA microarray, is driven by the suprachiasmatic nucleus. Curr Biol. 2002;12(7):540–550. - PubMed
-
- Panda S, et al. Coordinated transcription of key pathways in the mouse by the circadian clock. Cell. 2002;109(3):307–320. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

