Transcription and imprinting dynamics in developing postnatal male germline stem cells
- PMID: 26545815
- PMCID: PMC4647563
- DOI: 10.1101/gad.261925.115
Transcription and imprinting dynamics in developing postnatal male germline stem cells
Abstract
Postnatal spermatogonial stem cells (SSCs) progress through proliferative and developmental stages to populate the testicular niche prior to productive spermatogenesis. To better understand, we conducted extensive genomic profiling at multiple postnatal stages on subpopulations enriched for particular markers (THY1, KIT, OCT4, ID4, or GFRa1). Overall, our profiles suggest three broad populations of spermatogonia in juveniles: (1) epithelial-like spermatogonia (THY1(+); high OCT4, ID4, and GFRa1), (2) more abundant mesenchymal-like spermatogonia (THY1(+); moderate OCT4 and ID4; high mesenchymal markers), and (3) (in older juveniles) abundant spermatogonia committing to gametogenesis (high KIT(+)). Epithelial-like spermatogonia displayed the expected imprinting patterns, but, surprisingly, mesenchymal-like spermatogonia lacked imprinting specifically at paternally imprinted loci but fully restored imprinting prior to puberty. Furthermore, mesenchymal-like spermatogonia also displayed developmentally linked DNA demethylation at meiotic genes and also at certain monoallelic neural genes (e.g., protocadherins and olfactory receptors). We also reveal novel candidate receptor-ligand networks involving SSCs and the developing niche. Taken together, neonates/juveniles contain heterogeneous epithelial-like or mesenchymal-like spermatogonial populations, with the latter displaying extensive DNA methylation/chromatin dynamics. We speculate that this plasticity helps SSCs proliferate and migrate within the developing seminiferous tubule, with proper niche interaction and membrane attachment reverting mesenchymal-like spermatogonial subtype cells back to an epithelial-like state with normal imprinting profiles.
Keywords: DNA methylation; germline; imprinting; monoallelic; spermatogonia; stem cells.
© 2015 Hammoud et al.; Published by Cold Spring Harbor Laboratory Press.
Figures


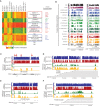


References
-
- Culty M. 2009. Gonocytes, the forgotten cells of the germ cell lineage. Birth Defects Res C Embryo Today 87: 1–26. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous
