Global analysis of biogenesis, stability and sub-cellular localization of lncRNAs mapping to intragenic regions of the human genome
- PMID: 26151857
- PMCID: PMC4615361
- DOI: 10.1080/15476286.2015.1062960
Global analysis of biogenesis, stability and sub-cellular localization of lncRNAs mapping to intragenic regions of the human genome
Abstract
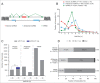
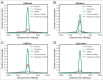
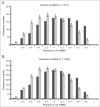
Long noncoding RNAs (lncRNAs) that map to intragenic regions of the human genome with the same (intronic lncRNAs) or opposite orientation (antisense lncRNAs) relative to protein-coding mRNAs have been largely dismissed from biochemical and functional characterization due to the belief that they are mRNA precursors, byproducts of RNA splicing or simply transcriptional noise. In this work, we used a custom microarray to investigate aspects of the biogenesis, processing, stability, evolutionary conservation, and cellular localization of ∼ 6,000 intronic lncRNAs and ∼ 10,000 antisense lncRNAs. Most intronic (2,903 of 3,427, 85%) and antisense lncRNAs (4,945 of 5,214, 95%) expressed in HeLa cells showed evidence of 5' cap modification, compatible with their transcription by RNAP II. Antisense lncRNAs (median t1/2 = 3.9 h) were significantly (p < 0.0001) more stable than mRNAs (median t1/2 = 3.2 h), whereas intronic lncRNAs (median t1/2 = 2.1 h) comprised a more heterogeneous class that included both stable (t1/2 > 3 h) and unstable (t1/2 < 1 h) transcripts. Intragenic lncRNAs display evidence of evolutionary conservation, have little/no coding potential and were ubiquitously detected in the cytoplasm. Notably, a fraction of the intronic and antisense lncRNAs (13 and 15%, respectively) were expressed from loci at which the corresponding host mRNA was not detected. The abundances of a subset of intronic/antisense lncRNAs were correlated (r ≥ |0.8|) with those of genes encoding proteins involved in cell division and DNA replication. Taken together, the findings of this study contribute novel biochemical and genomic information regarding intronic and antisense lncRNAs, supporting the notion that these classes include independently transcribed RNAs with potentials for exerting regulatory functions in the cell.
Keywords: Intronic lncRNAs; RNA stability; RNA subcellular localization; antisense lncRNAs; eukaryotic transcription.
Figures


References
-
- Bernstein BE, Birney E, Dunham I, Green ED, Gunter C, Snyder M. An integrated encyclopedia of DNA elements in the human genome. Nature 2012; 489:57-74; PMID:22955616; http://dx.doi.org/10.1038/nature11247 - DOI - PMC - PubMed
-
- Esteller M. Non-coding RNAs in human disease. Nat Rev Genet 2011; 12:861-74; PMID:22094949; http://dx.doi.org/10.1038/nrg3074 - DOI - PubMed
-
- Derrien T, Johnson R, Bussotti G, Tanzer A, Djebali S, Tilgner H, Guernec G, Martin D, Merkel A, Knowles DG, et al.. The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome Res 2012; 22:1775-89; PMID:22955988; http://dx.doi.org/10.1101/gr.132159.111 - DOI - PMC - PubMed
-
- Cabili MN, Trapnell C, Goff L, Koziol M, Tazon-Vega B, Regev A, Rinn JL. Integrative annotation of human large intergenic noncoding RNAs reveals global properties and specific subclasses. Genes Dev 2011; 25:1915-27; PMID:21890647; http://dx.doi.org/10.1101/gad.17446611 - DOI - PMC - PubMed
-
- Mercer TR, Dinger ME, Sunkin SM, Mehler MF, Mattick JS. Specific expression of long noncoding RNAs in the mouse brain. Proc Natl Acad Sci U S A 2008; 105:716-21; PMID:18184812; http://dx.doi.org/10.1073/pnas.0706729105 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous
