Hidden genetic diversity in the green alga Spirogyra (Zygnematophyceae, Streptophyta)
- PMID: 22655677
- PMCID: PMC3527229
- DOI: 10.1186/1471-2148-12-77
Hidden genetic diversity in the green alga Spirogyra (Zygnematophyceae, Streptophyta)
Abstract
Background: The unbranched filamentous green alga Spirogyra (Streptophyta, Zygnemataceae) is easily recognizable based on its vegetative morphology, which shows one to several spiral chloroplasts. This simple structure falsely points to a low genetic diversity: Spirogyra is commonly excluded from phylogenetic analyses because the genus is known as a long-branch taxon caused by a high evolutionary rate.
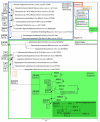
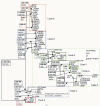
Results: We focused on this genetic diversity and sequenced 130 Spirogyra small subunit nuclear ribosomal DNA (SSU rDNA) strands of different origin. The resulting SSU rDNA sequences were used for phylogenetic analyses using complex evolutionary models (posterior probability, maximum likelihood, neighbor joining, and maximum parsimony methods). The sequences were between 1672 and 1779 nucleotides long. Sequence comparisons revealed 53 individual clones, but our results still support monophyly of the genus. Our data set did not contain a single slow-evolving taxon that would have been placed on a shorter branch compared to the remaining sequences. Out of 130 accessions analyzed, 72 showed a secondary loss of the 1506 group I intron, which formed a long-branched group within the genus. The phylogenetic relationship to the genus Spirotaenia was not resolved satisfactorily. The genetic distance within the genus Spirogyra exceeded the distances measured within any other genus of the remaining Zygnemataceae included in this study.
Conclusion: Overall, we define eight distinct clades of Spirogyra, one of them including the genus Sirogonium. A large number of non-homoplasious synapomorphies (NHS; 114 NHS in total) was found for Spirogyra (41 NHS) and for each clade (totaling 73 NHS). This emphasizes the high genetic diversity of this genus and the distance to the remaining Zygnematophyceae.
Figures

References
-
- Hoshaw RW, McCourt RM. The Zygnemataceae (Chlorophyta): A twenty-year update of research. Phycologia. 1988;27(4):511–548. doi: 10.2216/i0031-8884-27-4-511.1. - DOI
-
- Hainz R, Wöber C, Schagerl M. The relationship between Spirogyra (Zygnematophyceae, Streptophyta) filament type groups and environmental conditions in Central Europe. Aquat Bot. 2009;91(3):173–180. doi: 10.1016/j.aquabot.2009.05.004. - DOI
-
- Drummond CS, Hall J, Karol KG, Delwiche CF, McCourt RM. Phylogeny of Spirogyra and Sirogonium (Zygnematophyceae) based on rbcL sequence data. J Phycol. 2005;41(5):1055–1064. doi: 10.1111/j.1529-8817.2005.00130.x. - DOI
-
- Kadlubowska JZ. In: Süßwasserflora von Mitteleuropa, Chlorophyta VIII. Ettl H, Gerloff H, Heynig H, Mollenhauer D, editor. Gustav Fischer Verlag, Stuttgart, New York; 1984. Conjugatophyceae I - Zygnemales.
-
- Ashraf M, Godward MBE. Ultrastructure and chemistry of the zygospore wall of Spirogyra. Ann Bot. 1980;46:485–487.
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

