Comparative structural analysis of human DEAD-box RNA helicases
- PMID: 20941364
- PMCID: PMC2948006
- DOI: 10.1371/journal.pone.0012791
Comparative structural analysis of human DEAD-box RNA helicases
Abstract
DEAD-box RNA helicases play various, often critical, roles in all processes where RNAs are involved. Members of this family of proteins are linked to human disease, including cancer and viral infections. DEAD-box proteins contain two conserved domains that both contribute to RNA and ATP binding. Despite recent advances the molecular details of how these enzymes convert chemical energy into RNA remodeling is unknown. We present crystal structures of the isolated DEAD-domains of human DDX2A/eIF4A1, DDX2B/eIF4A2, DDX5, DDX10/DBP4, DDX18/myc-regulated DEAD-box protein, DDX20, DDX47, DDX52/ROK1, and DDX53/CAGE, and of the helicase domains of DDX25 and DDX41. Together with prior knowledge this enables a family-wide comparative structural analysis. We propose a general mechanism for opening of the RNA binding site. This analysis also provides insights into the diversity of DExD/H- proteins, with implications for understanding the functions of individual family members.
Conflict of interest statement
Figures

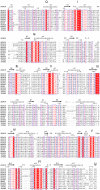




References
-
- Linder P, Tanner NK, Banroques J. From RNA helicases to RNPases. Trends Biochem Sci. 2001;26:339–341. - PubMed
-
- Schwer B. A new twist on RNA helicases: DExH/D box proteins as RNPases. Nat Struct Biol. 2001;8:113–116. - PubMed
-
- Cordin O, Banroques J, Tanner NK, Linder P. The DEAD-box protein family of RNA helicases. Gene. 2006;367:17–37. - PubMed
-
- Pyle AM. Translocation and unwinding mechanisms of RNA and DNA helicases. Annu Rev Biophys. 2008;37:317–336. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

