Structural basis for TetM-mediated tetracycline resistance
- PMID: 23027944
- PMCID: PMC3479509
- DOI: 10.1073/pnas.1208037109
Structural basis for TetM-mediated tetracycline resistance
Abstract
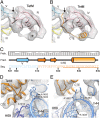
Ribosome protection proteins (RPPs) confer tetracycline resistance by binding to the ribosome and chasing the drug from its binding site. The current model for the mechanism of action of RPPs proposes that drug release is indirect and achieved via conformational changes within the drug-binding site induced upon binding of the RPP to the ribosome. Here we report a cryo-EM structure of the RPP TetM in complex with the 70S ribosome at 7.2-Å resolution. The structure reveals the contacts of TetM with the ribosome, including interaction between the conserved and functionally critical C-terminal extension of TetM and the decoding center of the small subunit. Moreover, we observe direct interaction between domain IV of TetM and the tetracycline binding site and identify residues critical for conferring tetracycline resistance. A model is presented whereby TetM directly dislodges tetracycline to confer resistance.
Conflict of interest statement
The authors declare no conflict of interest.
Figures




References
-
- Wilson DN. The A-Z of bacterial translation inhibitors. Crit Rev Biochem Mol Biol. 2009;44(6):393–433. - PubMed
-
- Brodersen DE, et al. The structural basis for the action of the antibiotics tetracycline, pactamycin, and hygromycin B on the 30S ribosomal subunit. Cell. 2000;103(7):1143–1154. - PubMed
-
- Roberts MC. Update on acquired tetracycline resistance genes. FEMS Microbiol Lett. 2005;245(2):195–203. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Medical

