Drug resistance against HCV NS3/4A inhibitors is defined by the balance of substrate recognition versus inhibitor binding
- PMID: 21084633
- PMCID: PMC3000259
- DOI: 10.1073/pnas.1006370107
Drug resistance against HCV NS3/4A inhibitors is defined by the balance of substrate recognition versus inhibitor binding
Abstract
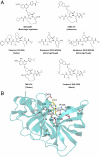
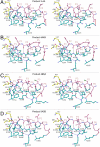
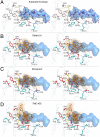
Hepatitis C virus infects an estimated 180 million people worldwide, prompting enormous efforts to develop inhibitors targeting the essential NS3/4A protease. Resistance against the most promising protease inhibitors, telaprevir, boceprevir, and ITMN-191, has emerged in clinical trials. In this study, crystal structures of the NS3/4A protease domain reveal that viral substrates bind to the protease active site in a conserved manner defining a consensus volume, or substrate envelope. Mutations that confer the most severe resistance in the clinic occur where the inhibitors protrude from the substrate envelope, as these changes selectively weaken inhibitor binding without compromising the binding of substrates. These findings suggest a general model for predicting the susceptibility of protease inhibitors to resistance: drugs designed to fit within the substrate envelope will be less susceptible to resistance, as mutations affecting inhibitor binding would simultaneously interfere with the recognition of viral substrates.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
Similar articles
-
The competitive binding between inhibitors and substrates of HCV NS3/4A protease: a general mechanism of drug resistance.Antiviral Res. 2014 Mar;103:60-70. doi: 10.1016/j.antiviral.2014.01.010. Epub 2014 Jan 23. Antiviral Res. 2014. PMID: 24462696
-
Understanding the structural and energetic basis of inhibitor and substrate bound to the full-length NS3/4A: insights from molecular dynamics simulation, binding free energy calculation and network analysis.Mol Biosyst. 2012 Oct;8(10):2753-65. doi: 10.1039/c2mb25157d. Mol Biosyst. 2012. PMID: 22833015
-
Computational study on the drug resistance mechanism against HCV NS3/4A protease inhibitors vaniprevir and MK-5172 by the combination use of molecular dynamics simulation, residue interaction network, and substrate envelope analysis.J Chem Inf Model. 2014 Feb 24;54(2):621-33. doi: 10.1021/ci400060j. Epub 2013 Jun 28. J Chem Inf Model. 2014. PMID: 23745769
-
Resistance outside the substrate envelope: hepatitis C NS3/4A protease inhibitors.Crit Rev Biochem Mol Biol. 2019 Feb;54(1):11-26. doi: 10.1080/10409238.2019.1568962. Epub 2019 Mar 1. Crit Rev Biochem Mol Biol. 2019. PMID: 30821513 Free PMC article. Review.
-
Discovery of small-molecule inhibitors of HCV NS3-4A protease as potential therapeutic agents against HCV infection.Curr Med Chem. 2005;12(20):2317-42. doi: 10.2174/0929867054864769. Curr Med Chem. 2005. PMID: 16181135 Review.
Cited by
-
A macrocyclic HCV NS3/4A protease inhibitor interacts with protease and helicase residues in the complex with its full-length target.Proc Natl Acad Sci U S A. 2011 Dec 27;108(52):21052-6. doi: 10.1073/pnas.1110534108. Epub 2011 Dec 12. Proc Natl Acad Sci U S A. 2011. PMID: 22160684 Free PMC article.
-
Coronaviruses resistant to a 3C-like protease inhibitor are attenuated for replication and pathogenesis, revealing a low genetic barrier but high fitness cost of resistance.J Virol. 2014 Oct;88(20):11886-98. doi: 10.1128/JVI.01528-14. Epub 2014 Aug 6. J Virol. 2014. PMID: 25100843 Free PMC article.
-
Hepatitis C Virus NS3/4A Protease Inhibitors Incorporating Flexible P2 Quinoxalines Target Drug Resistant Viral Variants.J Med Chem. 2017 Jul 13;60(13):5699-5716. doi: 10.1021/acs.jmedchem.7b00426. Epub 2017 Jun 19. J Med Chem. 2017. PMID: 28594175 Free PMC article.
-
Pharmacophore-Assisted Covalent Docking Identifies a Potential Covalent Inhibitor for Drug-Resistant Genotype 3 Variants of Hepatitis C Viral NS3/4A Serine Protease.Viruses. 2024 Aug 3;16(8):1250. doi: 10.3390/v16081250. Viruses. 2024. PMID: 39205224 Free PMC article.
-
Significant reduction in errors associated with nonbonded contacts in protein crystal structures: automated all-atom refinement with PrimeX.Acta Crystallogr D Biol Crystallogr. 2012 Aug;68(Pt 8):935-52. doi: 10.1107/S0907444912017453. Epub 2012 Jul 17. Acta Crystallogr D Biol Crystallogr. 2012. PMID: 22868759 Free PMC article.
References
-
- World Health Organization. . Vaccine research: hepatitis C. In: Barnes E, editor. Hepatitis C Virus: Disease Burden. 2010. [March 15, 2010]. Available at http://www.who.int/vaccine_research/diseases/viral_cancers/en/index2.html.
-
- Major ME, Feinstone SM. The molecular virology of hepatitis C. Hepatology. 1997;25:1527–1538. - PubMed
-
- Qureshi SA. Hepatitis C virus—biology, host evasion strategies, and promising new therapies on the horizon. Med Res Rev. 2007;27:353–373. - PubMed
-
- Paolucci S, et al. Analysis of HIV drug-resistant quasispecies in plasma, peripheral blood mononuclear cells and viral isolates from treatment-naive and HAART patients. J Med Virol. 2001;65:207–217. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

