Crystal structure of PhnZ in complex with substrate reveals a di-iron oxygenase mechanism for catabolism of organophosphonates
- PMID: 24706911
- PMCID: PMC3986159
- DOI: 10.1073/pnas.1320039111
Crystal structure of PhnZ in complex with substrate reveals a di-iron oxygenase mechanism for catabolism of organophosphonates
Erratum in
- Proc Natl Acad Sci U S A. 2014 Jun 3;111(22)8311
Abstract
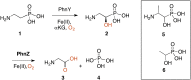
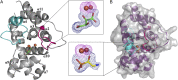
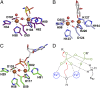
The enzymes PhnY and PhnZ comprise an oxidative catabolic pathway that enables marine bacteria to use 2-aminoethylphosphonic acid as a source of inorganic phosphate. PhnZ is notable for catalyzing the oxidative cleavage of a carbon-phosphorus bond using Fe(II) and dioxygen, despite belonging to a large family of hydrolytic enzymes, the HD-phosphohydrolase superfamily. We have determined high-resolution structures of PhnZ bound to its substrate, (R)-2-amino-1-hydroxyethylphosphonate (2.1 Å), and a buffer additive, l-tartrate (1.7 Å). The structures reveal PhnZ to have an active site containing two Fe ions coordinated by four histidines and two aspartates that is strikingly similar to the carbon-carbon bond cleaving enzyme, myo-inositol-oxygenase. The exception is Y24, which forms a transient ligand interaction at the dioxygen binding site of Fe2. Site-directed mutagenesis and kinetic analysis with substrate analogs revealed the roles of key active site residues. A fifth histidine that is conserved in the PhnZ subclade, H62, specifically interacts with the substrate 1-hydroxyl. The structures also revealed that Y24 and E27 mediate a unique induced-fit mechanism whereby E27 specifically recognizes the 2-amino group of the bound substrate and toggles the release of Y24 from the active site, thereby creating space for molecular oxygen to bind to Fe2. Structural comparisons of PhnZ reveal an evolutionary connection between Fe(II)-dependent hydrolysis of phosphate esters and oxidative carbon-phosphorus or carbon-carbon bond cleavage, thus uniting the diverse chemistries that are found in the HD superfamily.
Keywords: C–H bond activation; C–P bond cleavage; nonheme iron-dependent oxygenase; phosphonate.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

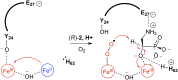
Similar articles
-
C-H Bond Cleavage Is Rate-Limiting for Oxidative C-P Bond Cleavage by the Mixed Valence Diiron-Dependent Oxygenase PhnZ.Biochemistry. 2019 Dec 31;58(52):5271-5280. doi: 10.1021/acs.biochem.9b00145. Epub 2019 May 10. Biochemistry. 2019. PMID: 31046250 Free PMC article.
-
Organophosphonate-degrading PhnZ reveals an emerging family of HD domain mixed-valent diiron oxygenases.Proc Natl Acad Sci U S A. 2013 Nov 19;110(47):18874-9. doi: 10.1073/pnas.1315927110. Epub 2013 Nov 6. Proc Natl Acad Sci U S A. 2013. PMID: 24198335 Free PMC article.
-
PhnY and PhnZ comprise a new oxidative pathway for enzymatic cleavage of a carbon-phosphorus bond.J Am Chem Soc. 2012 May 23;134(20):8364-7. doi: 10.1021/ja302072f. Epub 2012 May 11. J Am Chem Soc. 2012. PMID: 22564006
-
Phosphonate biosynthesis and catabolism: a treasure trove of unusual enzymology.Curr Opin Chem Biol. 2013 Aug;17(4):580-8. doi: 10.1016/j.cbpa.2013.06.018. Epub 2013 Jul 17. Curr Opin Chem Biol. 2013. PMID: 23870698 Free PMC article. Review.
-
Variations of the 2-His-1-carboxylate theme in mononuclear non-heme FeII oxygenases.Chembiochem. 2006 Oct;7(10):1536-48. doi: 10.1002/cbic.200600152. Chembiochem. 2006. PMID: 16858718 Review.
Cited by
-
Structure of Ddi2, a highly inducible detoxifying metalloenzyme from Saccharomyces cerevisiae.J Biol Chem. 2019 Jul 5;294(27):10674-10685. doi: 10.1074/jbc.RA118.006394. Epub 2019 May 31. J Biol Chem. 2019. PMID: 31152065 Free PMC article.
-
Transporter characterisation reveals aminoethylphosphonate mineralisation as a key step in the marine phosphorus redox cycle.Nat Commun. 2021 Jul 27;12(1):4554. doi: 10.1038/s41467-021-24646-z. Nat Commun. 2021. PMID: 34315891 Free PMC article.
-
Global and seasonal variation of marine phosphonate metabolism.ISME J. 2022 Sep;16(9):2198-2212. doi: 10.1038/s41396-022-01266-z. Epub 2022 Jun 23. ISME J. 2022. PMID: 35739297 Free PMC article.
-
Crystal structures of SAMHD1 inhibitor complexes reveal the mechanism of water-mediated dNTP hydrolysis.Nat Commun. 2020 Jun 23;11(1):3165. doi: 10.1038/s41467-020-16983-2. Nat Commun. 2020. PMID: 32576829 Free PMC article.
-
The Microbial Degradation of Natural and Anthropogenic Phosphonates.Molecules. 2023 Sep 29;28(19):6863. doi: 10.3390/molecules28196863. Molecules. 2023. PMID: 37836707 Free PMC article. Review.
References
-
- Dyhrman ST, Benitez-Nelson CR, Orchard ED, Haley ST, Pellechia PJ. A microbial source of phosphonates in oligotrophic marine systems. Nat Geosci. 2009;2(10):696–699.
-
- Dyhrman ST, et al. Phosphonate utilization by the globally important marine diazotroph Trichodesmium. Nature. 2006;439(7072):68–71. - PubMed
-
- Hudson JJ, Taylor WD, Schindler DW. Phosphate concentrations in lakes. Nature. 2000;406(6791):54–56. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

