Structural mechanism of glutamate receptor activation and desensitization
- PMID: 25119039
- PMCID: PMC4199900
- DOI: 10.1038/nature13603
Structural mechanism of glutamate receptor activation and desensitization
Abstract
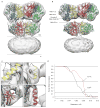
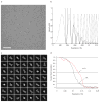
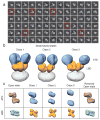
Ionotropic glutamate receptors are ligand-gated ion channels that mediate excitatory synaptic transmission in the vertebrate brain. To gain a better understanding of how structural changes gate ion flux across the membrane, we trapped rat AMPA (α-amino-3-hydroxy-5-methyl-4-isoxazole propionic acid) and kainate receptor subtypes in their major functional states and analysed the resulting structures using cryo-electron microscopy. We show that transition to the active state involves a 'corkscrew' motion of the receptor assembly, driven by closure of the ligand-binding domain. Desensitization is accompanied by disruption of the amino-terminal domain tetramer in AMPA, but not kainate, receptors with a two-fold to four-fold symmetry transition in the ligand-binding domains in both subtypes. The 7.6 Å structure of a desensitized kainate receptor shows how these changes accommodate channel closing. These findings integrate previous physiological, biochemical and structural analyses of glutamate receptors and provide a molecular explanation for key steps in receptor gating.
Conflict of interest statement
The authors declare no competing financial interests.
Figures











References
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

