Structure, function, and allosteric modulation of NMDA receptors
- PMID: 30037851
- PMCID: PMC6080888
- DOI: 10.1085/jgp.201812032
Structure, function, and allosteric modulation of NMDA receptors
Abstract
NMDA-type glutamate receptors are ligand-gated ion channels that mediate a Ca2+-permeable component of excitatory neurotransmission in the central nervous system (CNS). They are expressed throughout the CNS and play key physiological roles in synaptic function, such as synaptic plasticity, learning, and memory. NMDA receptors are also implicated in the pathophysiology of several CNS disorders and more recently have been identified as a locus for disease-associated genomic variation. NMDA receptors exist as a diverse array of subtypes formed by variation in assembly of seven subunits (GluN1, GluN2A-D, and GluN3A-B) into tetrameric receptor complexes. These NMDA receptor subtypes show unique structural features that account for their distinct functional and pharmacological properties allowing precise tuning of their physiological roles. Here, we review the relationship between NMDA receptor structure and function with an emphasis on emerging atomic resolution structures, which begin to explain unique features of this receptor.
© 2018 Hansen et al.
Figures


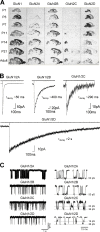


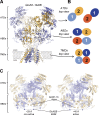



References
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

