Mechanisms of recognition of amyloid-β (Aβ) monomer, oligomer, and fibril by homologous antibodies
- PMID: 28924036
- PMCID: PMC5672054
- DOI: 10.1074/jbc.M117.801514
Mechanisms of recognition of amyloid-β (Aβ) monomer, oligomer, and fibril by homologous antibodies
Abstract
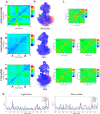
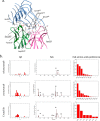
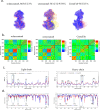
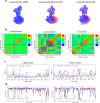
Alzheimer's disease is one of the most devastating neurodegenerative diseases without effective therapies. Immunotherapy is a promising approach, but amyloid antibody structural information is limited. Here we simulate the recognition of monomeric, oligomeric, and fibril amyloid-β (Aβ) by three homologous antibodies (solanezumab, crenezumab, and their chimera, CreneFab). Solanezumab only binds the monomer, whereas crenezumab and CreneFab can recognize different oligomerization states; however, the structural basis for this observation is not understood. We successfully identified stable complexes of crenezumab with Aβ pentamer (oligomer model) and 16-mer (fibril model). It is noteworthy that solanezumab targets Aβ residues 16-26 preferentially in the monomeric state; conversely, crenezumab consistently targets residues 13-16 in different oligomeric states. Unlike the buried monomeric peptide in solanezumab's complementarity-determining region, crenezumab binds the oligomer's lateral and edge residues. Surprisingly, crenezumab's complementarity-determining region loops can effectively bind the Aβ fibril lateral surface around the same 13-16 region. The constant domain influences antigen recognition through entropy redistribution. Different constant domain residues in solanezumab/crenezumab/chimera influence the binding of Aβ aggregates. Collectively, we provide molecular insight into the recognition mechanisms facilitating antibody design.
Keywords: Alzheimer disease; amyloid; amyloid-beta (AB); antibody; immunotherapy; protein dynamic.
Conflict of interest statement
The authors declare that they have no conflicts of interest with the contents of this article
Figures






References
-
- Ballard C., Gauthier S., Corbett A., Brayne C., Aarsland D., and Jones E. (2011) Alzheimer's disease. Lancet 377, 1019–1031 - PubMed
-
- Hardy J. A., and Higgins G. A. (1992) Alzheimer's disease: the amyloid cascade hypothesis. Science 256, 184–185 - PubMed
-
- De Genst E., Messer A., and Dobson C. M. (2014) Antibodies and protein misfolding: from structural research tools to therapeutic strategies. Biochim. Biophys. Acta 1844, 1907–1919 - PubMed
-
- Herrmann A., and Spires-Jones T. (2015) Clearing the way for tau immunotherapy in Alzheimer's disease. J. Neurochem. 132, 1–4 - PubMed
-
- Hardy J., Bogdanovic N., Winblad B., Portelius E., Andreasen N., Cedazo-Minguez A., and Zetterberg H. (2014) Pathways to Alzheimer's disease. J. Intern. Med. 275, 296–303 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

